Project A.11
- PhD student: Neel Wagh
- Supervisor: Marion Silies
- Further TAC-members: Carlotta Martelli , Martin Heine
- Research Group
Most animals rely on their visual system for many daily tasks. Effective processing of visual sensory information is fundamental to life across most animal species. We use the geneticist’s favorite model, Drosophila, to understand the genetic and gene regulatory mechanisms underlying neuronal properties and how they shape stable visual processing. Using genetic analysis in combination with in-vivo 2photon calcium imaging and complementary behavioral assays we investigate the neural computations fundamental to visually guided behavior.
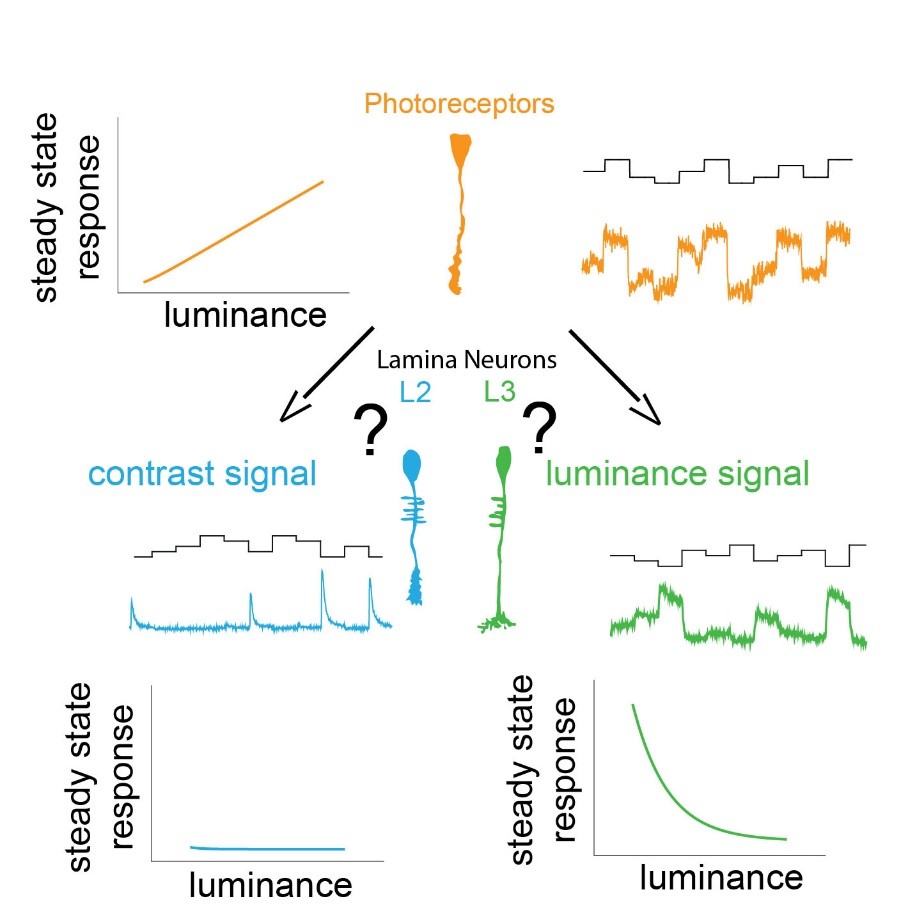
We use our eyes constantly to gather information about our surroundings to perform tasks throughout the day. We need to sample our visual environment and build visual representation updating in time to guide appropriate behavior. A key feature of the visual system is to extract contrast, which is the basis for all visual computations. The stable computation of contrast requires two circuit components: a pathway that responds to changes in the visual scene and a luminance-sensitive pathway that provides a fast luminance gain when background luminance quickly changes. The early visual processing in lamina shows functional similarities between many insects like the blowfly, dragonfly, butterfly, worker bees. The lamina of blowfly and dragonfly are anatomically different yet show similar function implying convergent evolution. Insects adapted to dim light conditions like cockroaches and moths suggest presence of luminance gain for navigation in such light conditions. Thus, understanding the principles behind the two circuit components of stable contrast computation is essential. We can decipher the underlying mechanisms using resourceful and genetically amenable Drosophila melanogaster.
In Drosophila melanogaster, these two pathways diverge downstream of the same histaminergic photoreceptor input in two first-order interneurons: change-sensitive L2 and luminance-sensitive L3 neurons. How these two neurons obtain such fundamentally different properties despite receiving the same synaptic input is unknown.
These differences in physiological properties may be due to cell-autonomous mechanisms and/or circuit dependent. Initial findings reveal that the L3-specific transcription factor dFezf regulates L3’s luminance sensitivity. To decipher downstream players of this regulation, we will focus on genes that are transcriptionally regulated by dFezf, specifically ion channels and neurotransmitter receptors. For the genetic regulation of neuronal properties in Drosophila melanogaster, I will use a reverse genetics approach and pharmacology to assay the functional role of these players in the modulation of L2 vs L3 responses. Using in vivo 2-photon calcium imaging, I will specifically record calcium signals to investigate physiological properties of L2 and L3. I will complement these with behavioral assays to understand the functional, behavioral consequences of the genetic manipulations. Together, these experiments will reveal a comprehensive understanding of a critical visual processing step.